User Tools
Sidebar
This is an old revision of the document!
Table of Contents
Tools
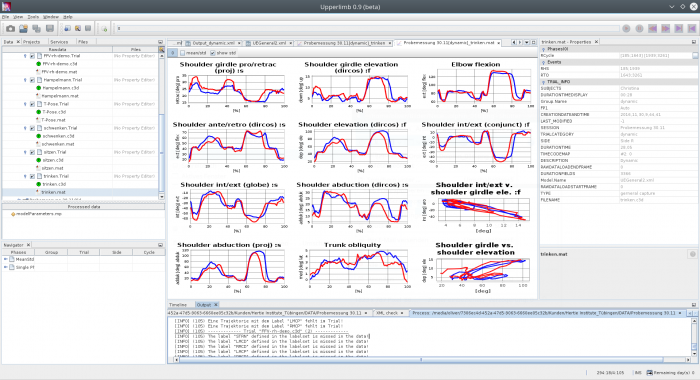
Plot Sheet
A “Plostsheet” component shows diagrams organized in rows and columns of a table in one or several pages. The count of diagrams, descriptions and the variables to show are configured in a specific xml-file saved in the “SheetsDefs” subfolder of a project definition. Very complex definitions are possible to show different sets of reference data behind the curves, to show markers e.g. an upright line at special time position or to compare different groups of patients.
Configuration
The main structure of the xml-configuration file looks as follow:
<Sheet name=""> <Pages> <Page rows="" columns=""> <Diagram name="" .../> </Page> </Pages> </Sheet>
Available features:
Save Mean+Std-Deviation of selected data
In the Plotsheets tab context menu, execute “Save Mean/Std”. This saves a file with the file suffix “.d3d” in the “output” subfolder of the opened main project. The filename includes the patient id (parent folder name) and delimited with “_” the name of the motion (sub folder name).
The structure of this file is
<subject>/<session>/<group>/<phase>/<original trial name>_<phase number> <subject>/<session>/<group>/<phase>/<original trial name>_<phase number> <subject>/<session>/<group>/<phase>/meanStd ... <group>/meanStd
That means that timeseries of time normalised phases and its mean+std are saved. The files can be used to create a reference data set as it is shown as grey bands behind patient data.
Datamining
In the Plotsheets tab context menu, execute “Datamining”.
This extract scalar-parameters from the timeseries based on a specific xml-configuration and saves them into a spreadsheet file e.g. for Microsoft excel or Libre/Open-Office.
View 4D
The View4d tool is a highly configurable 3d viewer. Details are described in the View 4D platform documentation.
Dataminer
The datamining tool determines and extracts specific defined parameters (single values) from timeseries data, e.g. maximum, minimum or mean values of specific phases or at specific positions (events). Details are described in the Dataminer platform documentation.
Group mean data wizzard
This component can be used to create reference data sets to be shown as gray bands behind the plosts. Details are described in the Group Mean Data Wizzard platform documentation.